
TIM is a data model that captures a common set of concepts and relationships for biomedical research intended to facilitate and encourage data sharing and reuse. Its purpose is to enable researchers to find highly connected biomedical data in a federated search space and support interoperability among datasets.
Examples of searches include:
TIM is formally specified using OWL. Although the human-readable document is rather limited at this time, TIM is published both in human- and machine-readable forms. A GET request leverages HTTP Content Negotiation to return the appropriate format. The available serializations are available directly using the links below.
The Terra Interoperability Model (TIM) is currently under development. Our github repository is terra-interoperability-model. Both JSON format and Terra Data Repository schema implementations are anticipated.
Release 2.x is currently available.
The TerraCore imports two other model files, TerraDCAT_ap and TerraCoreValueSets; together these three files make up the Terra Interoperability Model (TIM).
Terra Interoperability Model
TerraCoreDataModel.ttl
TerraCoreValueSets.ttl
TerraDCAT-AP.ttl
References
TIM also imports subsets of several well-known ontologies, including
UBERON,
Ontology of Biomedical Investigations,
NCI Thesaurus OBO Edition,
EFO, and
Cell Ontology.
The referenced subsets can be directly accessed at the links below:
CL_subset,
EFO_subset,
NCBITaxon_Organisms_subset,
OBI_assay_subset,
OBI_core,
UBERON_subset
Please use the repository's Issue Tracker to share comments or concerns related to the data model. TIM is not currently open to public contribution; it is currently wholly managed by Broad’s DSP Data Modeling Team.
The following diagram illustrates selected key concepts and relationships to provide an overview of the model.
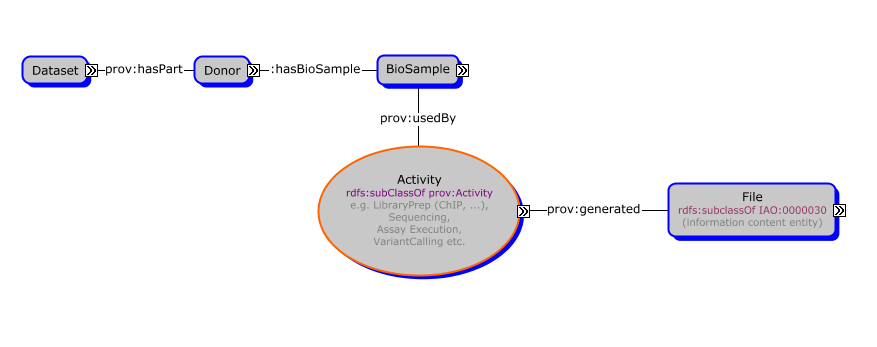
Prefixes are shorthand to reference concepts and properties in another namespace. The following prefixes are used in the illustration.
Diagram produced using Cmap from the Florida Institute for Human & Machine Cognition (IHMC).